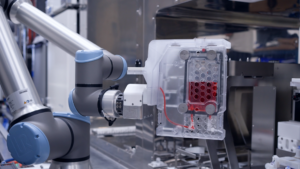
Roche Scales NVIDIA AI Factories Globally to Accelerate Drug Discovery, Diagnostic Solutions and Manufacturing Breakthroughs
Roche's new deployment spans more than 3,500 NVIDIA Blackwell GPUs across its worldwide operations and embedded across the entire value chain, massively scaling R&D productivity, next-generation diagnostics and manufacturing efficiencies....