Researchers are racing to discover the right drug molecule to treat COVID-19 — but the number of potential drug-like molecules out there is estimated to be an inconceivable 1060.
“Even if you hypothetically checked one molecule per second, it would take longer than the age of the universe to explore the entire chemical space,” said Shinya Yuki, co-founder and CEO of Tokyo-based startup Elix, Inc. “AI can efficiently explore huge search spaces to solve difficult problems, whether in drug discovery, materials development or a game like Go.”
Yuki’s company is using deep learning to accelerate drug discovery, building neural networks that predict the properties of molecules much faster than computer simulations can. To support COVID-19 research, the team is using AI to find drugs that are FDA-approved or in clinical trials that could be repurposed to treat the coronavirus.
“Developing a new drug from scratch is a years-long process, which is unwanted especially in this pandemic situation,” Yuki said. “Speed is critical, and drug-repurposing can help identify candidates with an existing clinical safety record, significantly reducing the time and cost of drug development.”
Elix recently published a paper on approved and clinical trial-stage drugs that its AI model flagged for potential COVID-19 treatments. Among the candidates selected by Elix’s AI tool was remdevisir, an antiviral drug that recently received emergency use authorization from the FDA for coronavirus cases.
A member of NVIDIA Inception, a program that helps startups get to market faster, Elix uses the NVIDIA DGX Station for training and inference of its deep learning algorithms. Yuki spoke about the company’s work in AI for drug discovery in the Inception Startup Showcase at GTC Digital, NVIDIA’s digital conference for developers and AI researchers.
Elix’s AI Fix for Drug Discovery
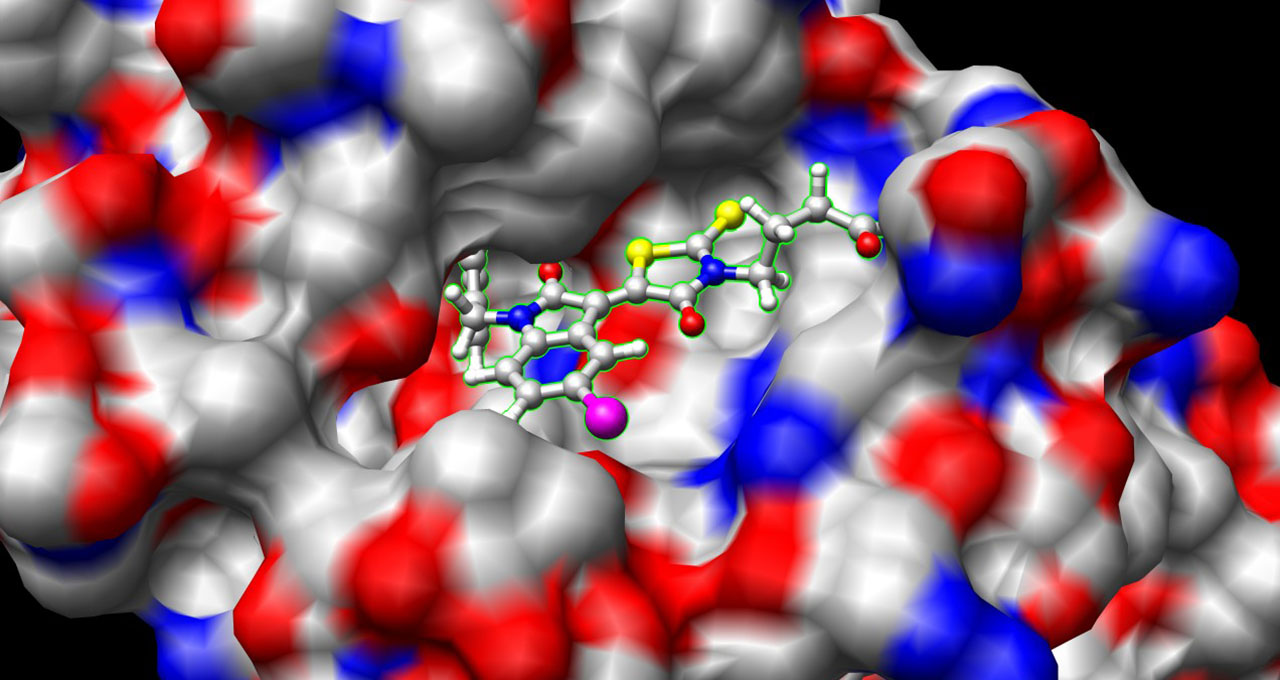
At the molecular level, a successful drug must have the perfect combination of shape, flexibility and interaction energies to bind to a target protein — like the spike proteins that cover the viral envelope of SARS-CoV-2, the virus that causes COVID-19.
A person gets infected with COVID-19 when these spike proteins attach to cells in the body, bringing the virus into the cells. An effective antiviral drug might interfere with this attachment process. For example, a promising drug molecule would bind with receptors on the spike proteins, preventing the virus from attaching to human cells.
To help researchers find the best drug for the job, Elix uses a variety of neural networks to rapidly narrow down the field of potential molecules. This allows researchers to reserve physical tests in the lab for a smaller subset of molecules that have a higher likelihood of solving the problem.
With predictive AI models, Yuki’s team can analyze a database of drug candidates to infer which have the right physical and chemical properties to treat a given disease. They also use generative models, which start from scratch to come up with promising molecular structures — some of which may not be found in nature.
That’s where a third neural network comes in, a retrosynthesis model that helps researchers figure out if the generated molecules can be synthesized in the lab.
Elix uses multiple NVIDIA DGX Station systems — GPU-powered AI workstations for data science development teams — to accelerate training and inference of these neural networks, achieving up to a 6x speedup using a single GPU for training versus a CPU.
Yuki says the acceleration is essential for the generative models, which would otherwise take a week or more to train until convergence, when the neural network reaches the lowest error rate possible. Each DGX Station has four NVIDIA V100 Tensor Core GPUs, enabling the Elix team to tackle bigger AI models and run multiple experiments at once.
“DGX Stations are basically supercomputers. We usually have several users working on the same machine at the same time,” he said. “We can not only train models faster, we can also run up to 15 experiments in parallel.”
The startup’s customers include pharmaceutical companies, research institutes and universities. Since molecular data is sensitive intellectual property for the pharma industry, most choose to run the AI models on their own on-prem servers.
Beyond drug discovery, Elix also uses AI for molecular design for material informatics, working with companies like tire- and rubber-manufacturer Bridgestone and RIKEN, Japan’s largest research institution. The company also develops computer vision models for autonomous driving and AI at the edge.
In one project, Yuki’s team worked with global chemical company Nippon Shokubai to generate a molecule that can be used as a blending material for ink, while posing a low risk of skin irritation.
Learn more about Elix in Yuki’s GTC Digital lightning talk. Visit our COVID page to explore how other startups are using AI and accelerated computing to fight the pandemic and stay up to date with our latest healthcare news.
Main image by Chaos, licensed from Wikimedia Commons under CC BY-SA 3.0.