Scientists and healthcare researchers — once limited by the number of samples that could be studied in a wet lab, or the quality of microscopes to peer into cells — are harnessing powerful computational tools to draw insights from an ever-growing trove of biological data.
Backing this digital biology revolution is a combination of high performance computing systems and domain-specific software frameworks.
Two supercomputers featured in the TOP500 ranking of most powerful systems announced today — NVIDIA’s healthcare-focused Cambridge-1 and biotechnology company Recursion’s BioHive-1 — are based on the NVIDIA DGX SuperPOD reference architecture.
And medical research institutions, pharmaceutical companies and biotech startups worldwide use NVIDIA Clara Parabricks, a suite of genomics libraries and reference applications, to power next-generation sequencing workflows.
Shanghai-based Mingma Biotechnology this month became the first research lab in China to launch Clara Parabricks Pipelines to support its work in precision medicine. This follows large-scale genomics initiatives rolled out in Thailand and Japan this year. And gene therapy startup Greffex recently adopted Parabricks Pipelines to accelerate its project to develop a universal influenza vaccine.
Identifying Genomic Insights for Population Studies
Parabricks Pipelines speed up DNA- and RNA-based projects by up to 50x on NVIDIA GPUs, enabling scientists to extract as much useful information as possible from the hundreds of terabytes of instrument data generated daily. This acceleration is especially powerful for public health institutions and research labs running population studies with tens of thousands of genomes to be analyzed.
Mingma Biotechnology adopted Parabricks Pipelines and NVIDIA T4 Tensor Core GPUs to accelerate its work in sequencing and multi-omics data analysis. The company provides medical institutions, pharmaceutical companies and researchers with genomic insights for disease research and drug development.
At the National Biobank of Thailand, an NVIDIA DGX A100 system is powering Genomics Thailand, an initiative to introduce genomic medicine as a common healthcare service in the country. The research institution is using Parabricks Pipelines to analyze genetic variations from 50,000 Thai volunteers’ whole genome sequencing data.
Pairing the DGX system with Parabricks Pipelines reduced the project’s whole genome data processing time by four months. Insights from this work will help researchers better analyze genetic variation specific to the Thai population.
And in Japan, the University of Tokyo’s Human Genome Center recently launched SHIROKANE, the country’s fastest supercomputer for life sciences. The DGX A100-powered system is running Parabricks Pipelines to sequence whole genomes of 92,000 patients, creating a database that is foundational to precision medicine efforts for cancer and intractable diseases.
Powering Clinical Sequencing, Drug Discovery
The Parabricks Pipelines suite of genetic tools can be configured to meet each laboratory’s specific needs. Researchers run Parabricks Pipelines workloads on NVIDIA GPU systems that range from desktop workstations to GPU-accelerated clouds and some of the world’s fastest supercomputers.
Within weeks of getting started with an HP data science workstation featuring NVIDIA RTX GPUs, Houston-based Greffex is using Parabricks Pipelines and NVIDIA Clara Discovery to advance its efforts to develop a universal flu vaccine.
The startup uses a combination of genomic sequencing, molecular dynamics tools and wet lab research to study how influenza strains evolve over time, and how these mutations impact vaccine efficacy.
To monitor changes in the flu, Greffex collects tens of thousands of flu genomes from around the world and runs massive sequence alignments on NVIDIA RTX 8000 GPUs to identify where the virus’s genetic code is changing. Running genomic workloads on GPUs is saving the company up to 13 hours per sample, while also enabling its team to rerun samples with different parameters to fine-tune the alignment results.
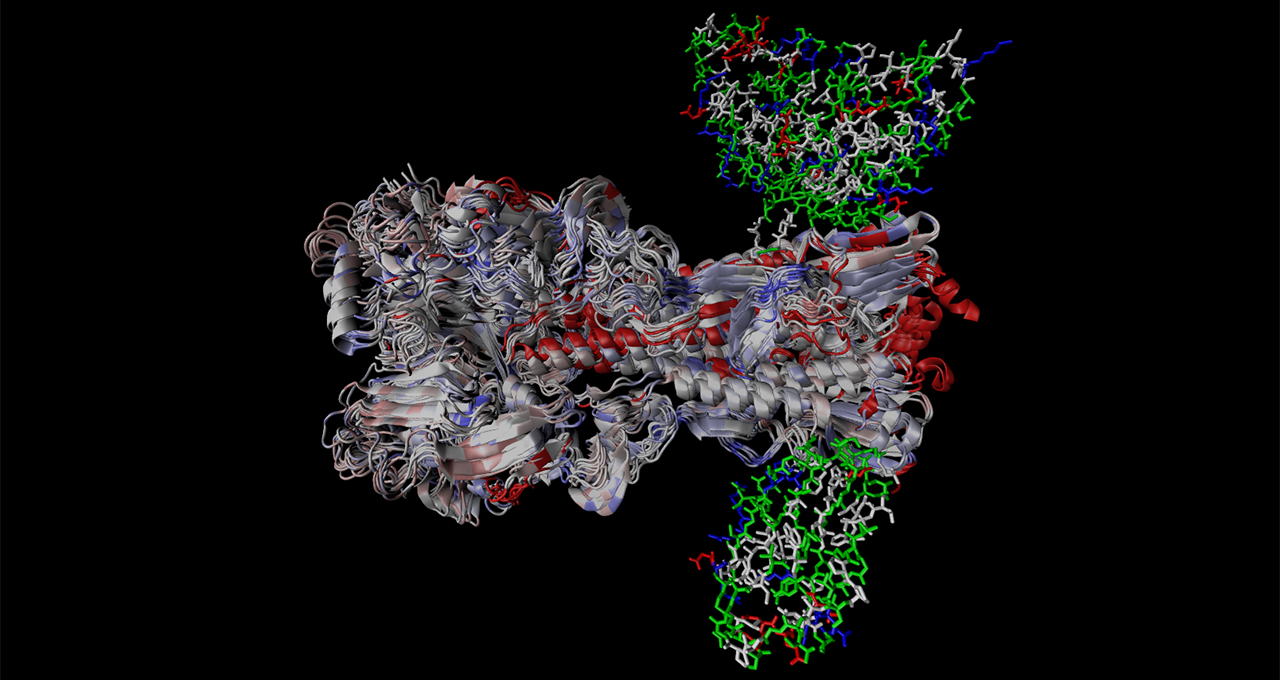
Once genetic variants are identified, Greffex scientists use molecular dynamics to visualize how these genetic changes alter the physical shape of the flu virus. They’re on the lookout for divergent mutations, where the flu virus may morph to a shape that doesn’t bind as well with vaccine-prompted antibodies.
“It’s a very lengthy and expensive process to optimize the protein structure for a vaccine that binds not just with the current flu strain, but a bunch of other strains,” said Daniel Preston, a bioinformatics scientist at Greffex. “With computational methods, we can get a sense of what will likely work before testing in real-world labs. It’s like using a scalpel versus using a hammer.”
About NVIDIA Clara Parabricks
NVIDIA Clara Parabricks brings GPU speedups to the Broad Institute’s industry-standard Genome Analysis Toolkit, as well as popular tools like Google’s DeepVariant genetic caller. Running on NVIDIA A100 Tensor Core GPUs, Parabricks accelerates secondary analysis times of a whole human genome to 23 minutes for DNA germline variant calling, compared to more than 20 hours on a CPU system.
Starting with DNA sequencing reads, Clara Parabricks Pipelines can align, sort, filter and call variants for germline and somatic variant detection, as well as support RNA-based applications. Germline variants are those inherited through an individual’s ancestry, while somatic mutations occur in human cells over a person’s lifetime and can lead to cancer.
Parabricks Pipelines version 3.6, releasing next month, will offer more tools for somatic variant calling — which gives researchers insights for precision oncology — and de novo germline variant calling, which informs research into complex diseases like autism.
The de novo germline variant calling pipeline was developed in collaboration with researchers from the Washington University School of Medicine. The technique reduces runtime to under an hour to parse genome data and identify novel variants within a family pedigree or parent-child trio.
Get started with NVIDIA Clara Parabricks Pipelines for accelerated genome analysis on NGC or on the AWS Marketplace.
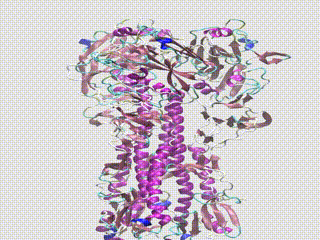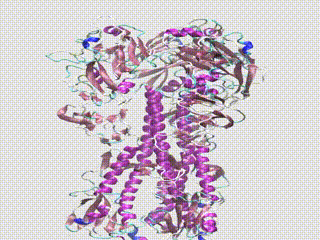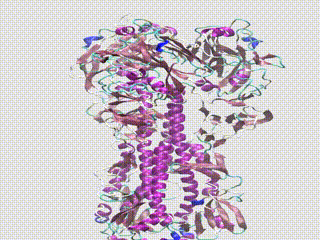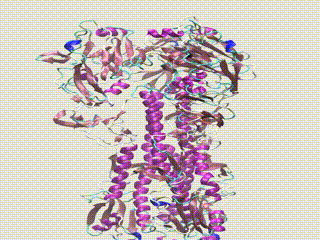
Main image shows a structural alignment of 17 H1 Hemagglutinin proteins spanning 102 years with simulated bound antibodies. Colored segments correspond to different types of mutations in the proteins. Image courtesy of Greffex, using models from the RCSB Protein Data Bank.